White Blood Cell Classification Challenge
2026
Abstract
Automatic classification of white blood cells from microscopic images is a major challenge for haematological diagnosis. This project addresses a 13-class classification problem from 28,901 training images with an extreme class imbalance (1183:1 ratio). We developed a classical ML pipeline based on 89 handcrafted features (morphological, colorimetric, textural) with dual cell/nucleus segmentation, then transitioned to deep learning with ConvNeXt-Tiny, Focal Loss, MixUp/CutMix, backbone freezing, and Test Time Augmentation. The iterative approach, guided by t-SNE embedding analysis and confusion matrix diagnostics, achieved a progression from 0.491 to 0.77 macro F1 on the Kaggle leaderboard. Ranked 20th out of 70+ students at Télécom Paris.
Classical ML: Handcrafted Features & Segmentation
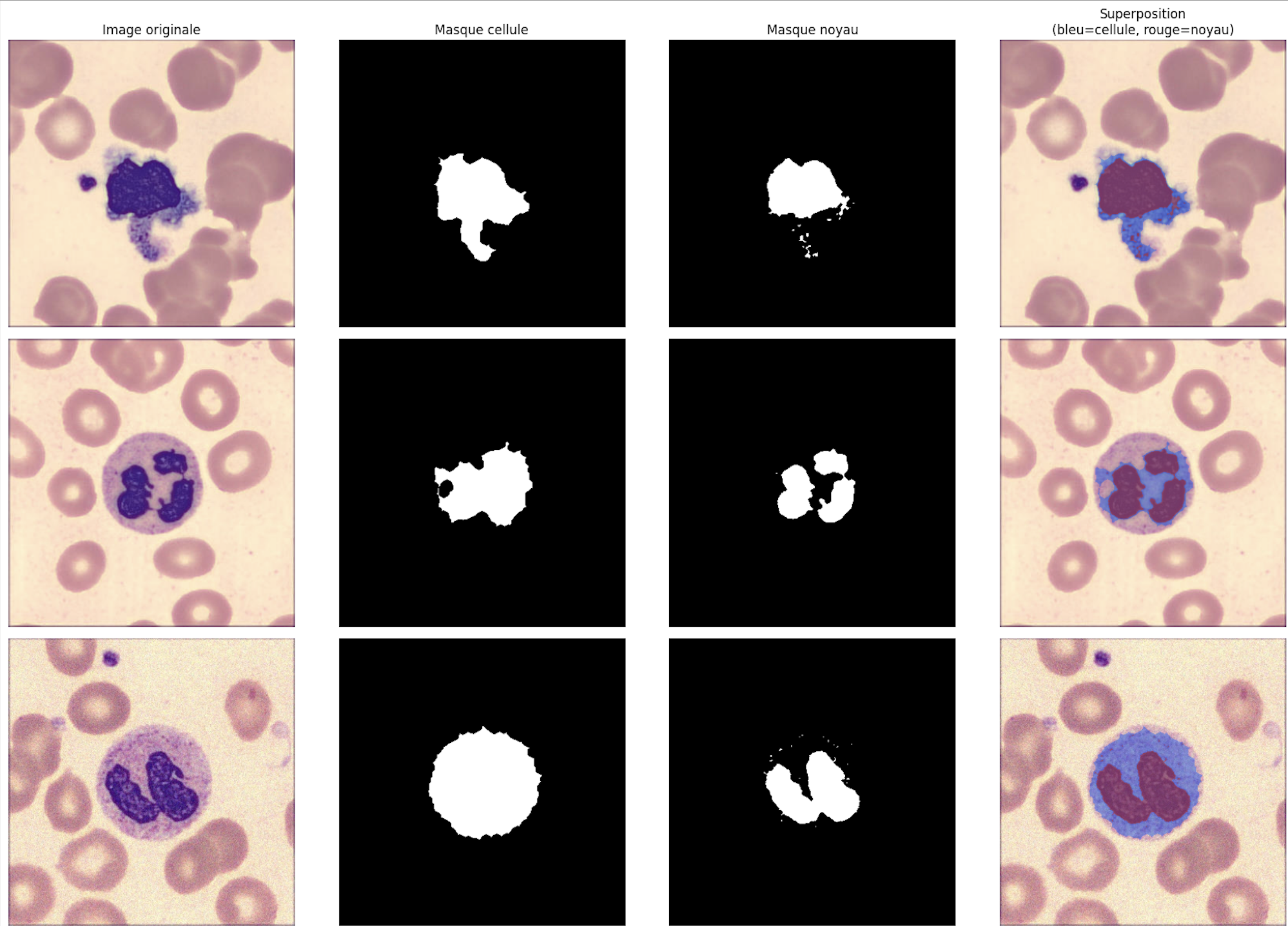
The first approach relies on a dual cell/nucleus segmentation using Otsu thresholding in CIELab colour space, followed by extraction of 89 handcrafted features spanning three categories: morphological (area, circularity, Hu moments, nucleus/cell ratio), colorimetric (RGB/HSV statistics per region: cell, nucleus, cytoplasm), and textural (GLCM properties over 4 angles + LBP histograms). Class imbalance is handled by SMOTE with k_neighbors=3 for ultra-rare classes and class_weight='balanced'. After GridSearchCV optimization, GradientBoosting achieves a macro F1 of 0.491. Confusion matrix analysis reveals that biologically similar classes (BNE/SNE, PLY/LY, MMY/MY/PMY) are systematically confused, highlighting the limits of handcrafted features for capturing fine morphological differences.
Deep Learning: From EfficientNet to ConvNeXt
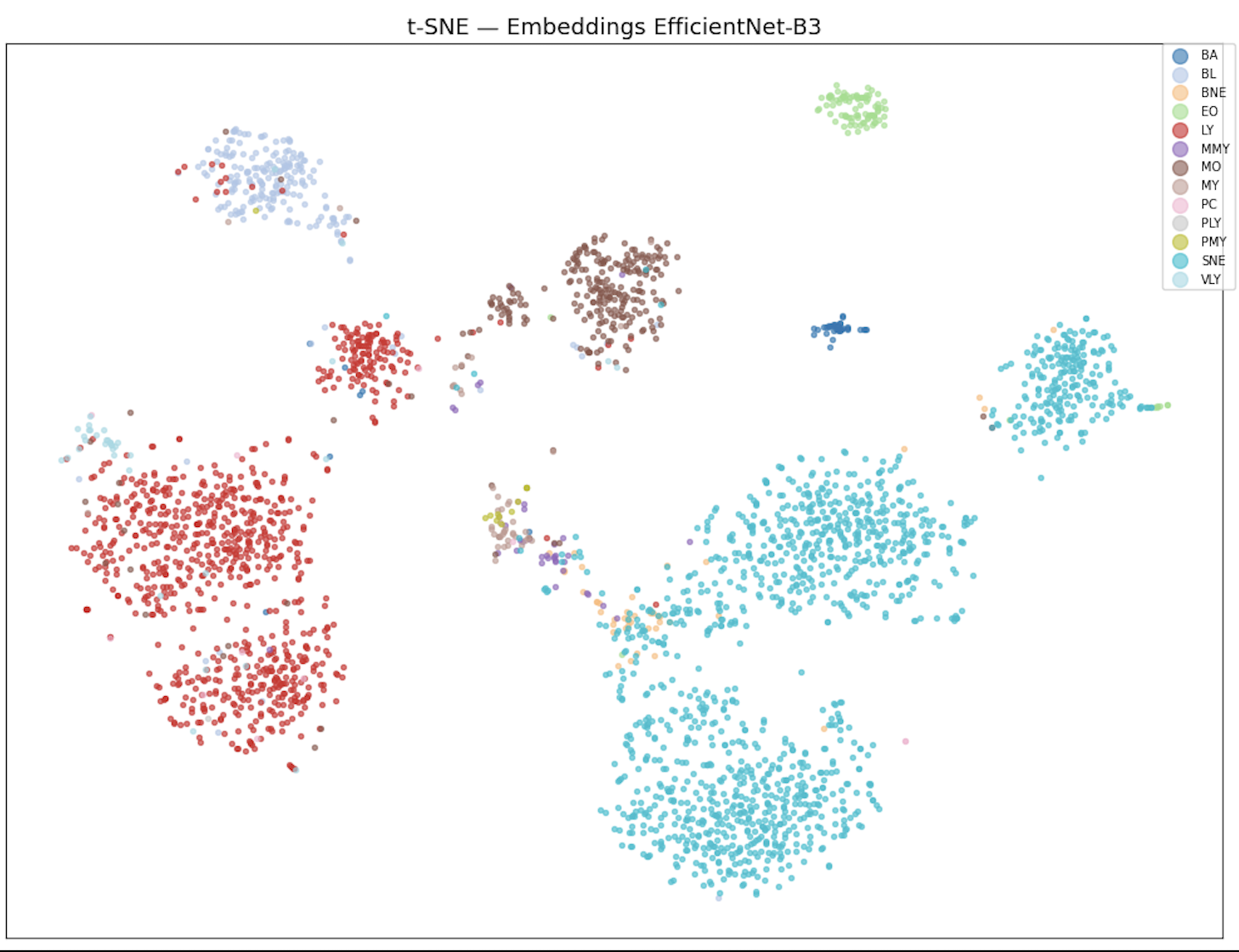
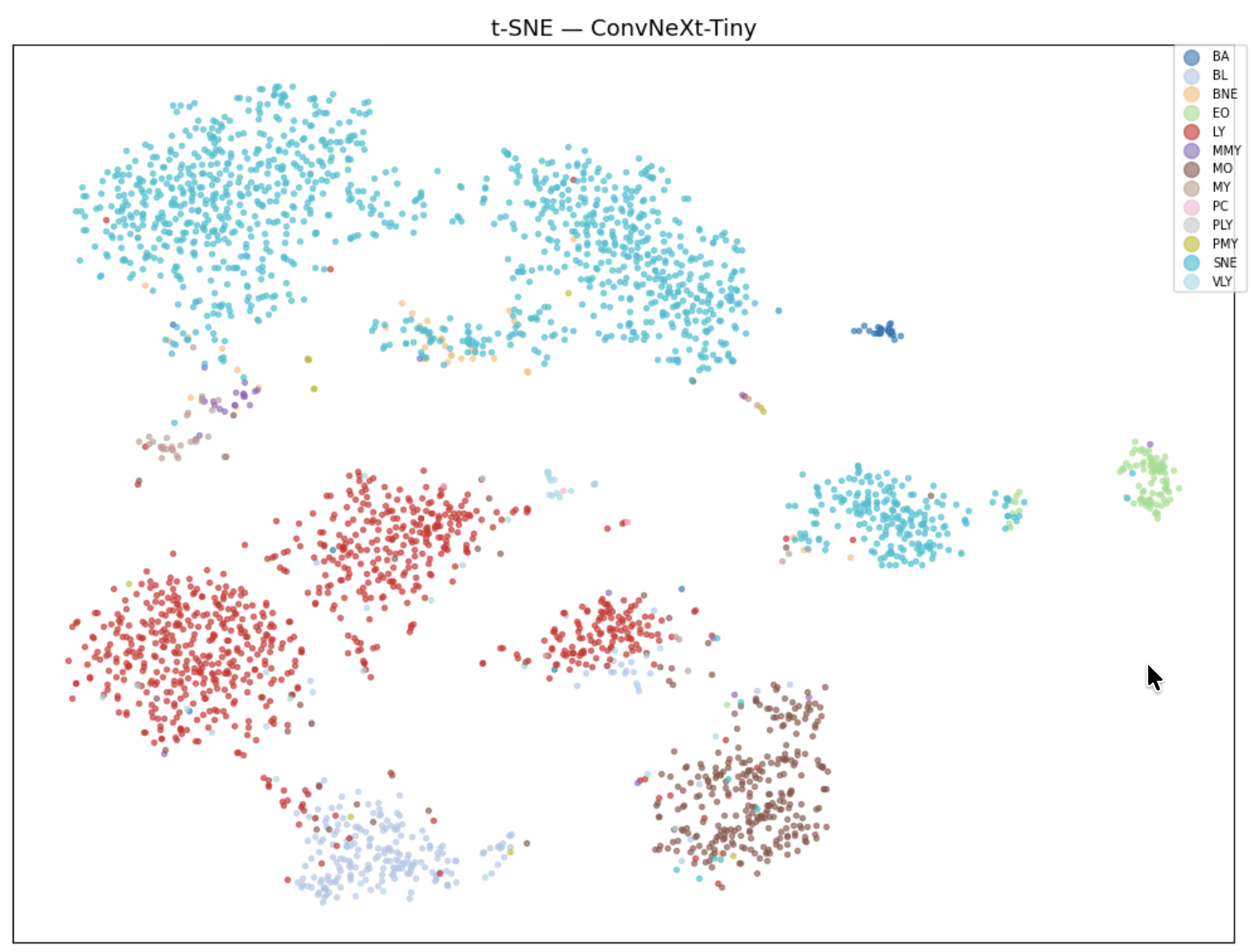
Transfer learning with EfficientNet-B3 pre-trained on ImageNet achieves 0.685 on the leaderboard, a +19 point jump over classical ML. However, t-SNE visualization of the embeddings reveals that rare classes (MMY, MY, PMY, PLY) form an indistinct central blob in feature space. This motivates the switch to ConvNeXt-Tiny with Focal Loss (gamma=2), which forces the network to focus on hard-to-classify examples. A CosineAnnealingWarmRestarts scheduler with T_0=15 over 60 epochs escapes local minima through periodic learning rate restarts, pushing the score to 0.747 with Test Time Augmentation (8 rounds, temperature scaling T=0.8).
Optimization: Diagnosing and Fixing Overfitting
A paradoxical observation drove the final optimization phase: adding MixUp improved validation F1 from 0.696 to 0.730, but the Kaggle score dropped from 0.747 to 0.728. This diagnosed overfitting on the validation split itself. Successive fixes included: backbone freezing for 5 epochs to prevent noisy gradient destruction of ImageNet features, CutMix alongside MixUp (50/50) for spatial regularization, weight decay increase from 5e-4 to 1e-2, and recalibration of class weights from log-dampened to 1/sqrt(n) with Focal Loss gamma reduced from 2.0 to 1.0. Each change was motivated by a specific diagnostic — not trial and error — yielding a final score of 0.77.
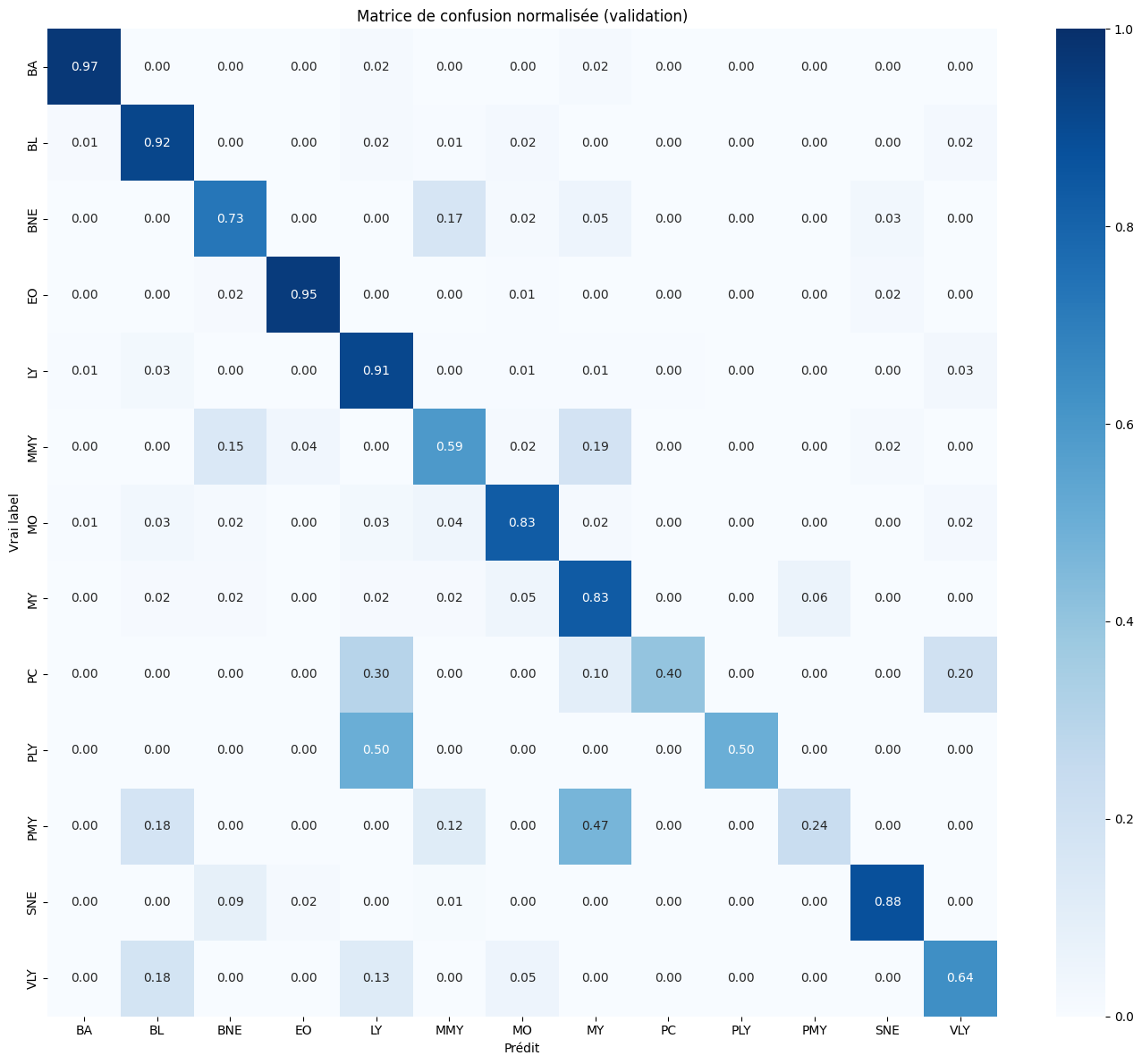
Results & Analysis
The optimized model achieves high recalls on dominant classes (BA: 0.97, EO: 0.95, LY: 0.91, SNE: 0.88) while rare classes remain challenging: PLY (0.50, confused with LY), PC (0.40), PMY (0.24, confused with MY). These residual confusions are biologically coherent — PLY is a lymphocyte precursor morphologically near-identical to LY, and the granulocytic lineage (MMY/MY/PMY) represents successive maturation stages with gradual morphological boundaries. With only 11 training images for PLY, no method can compensate for the fundamental lack of diversity. Overall progression: 0.491 (ML) → 0.685 (EfficientNet) → 0.747 (ConvNeXt + TTA) → 0.77 (optimized), a +28.3 point gain.